Canonical analysis of principal coordinates (CAP, left) ordination of baited video (above) and trap (below) data with corresponding strength and direction of Spearman correlation >0.35 of fish species shown as line vectors (

CAP summary. Canonical analysis of principal components (CAP). This is... | Download Scientific Diagram

CAP – Context, Analysis, Practice: A lesson planning model for language teacher education – Jason Anderson: Teacher educator & author

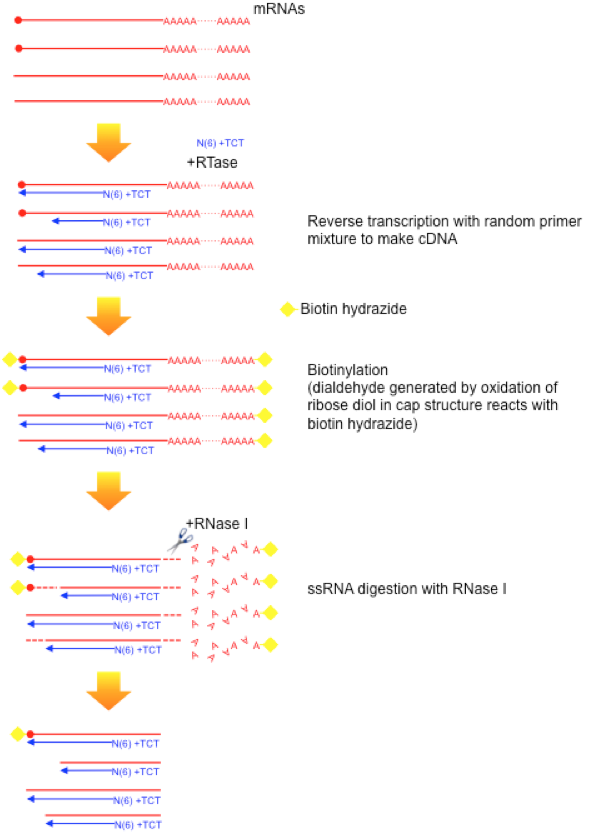
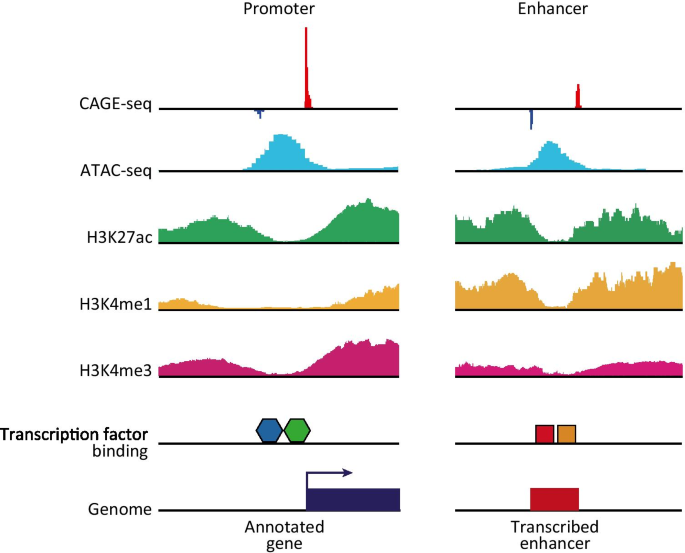
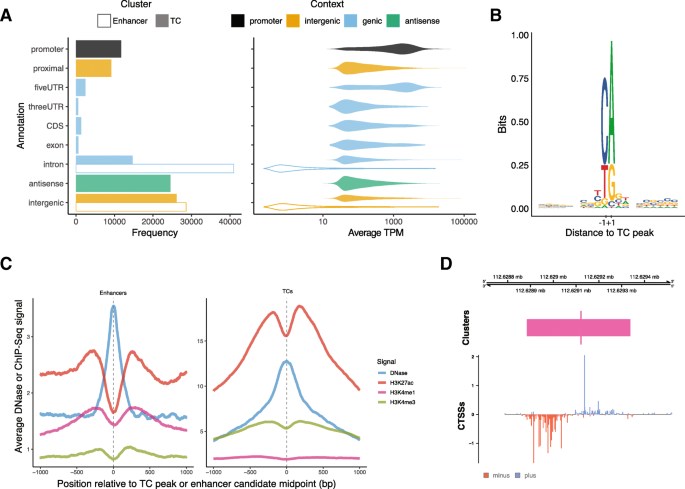
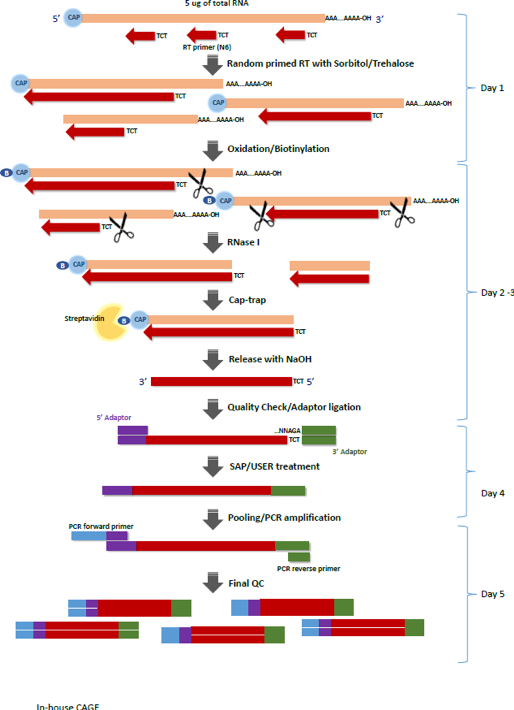
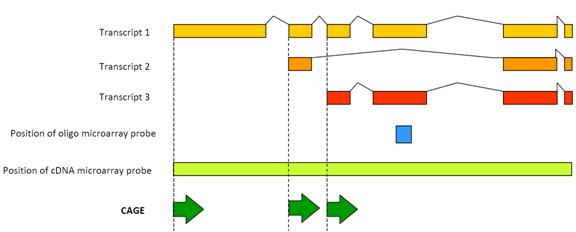
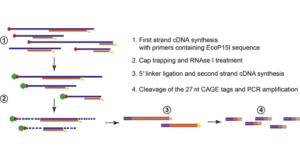
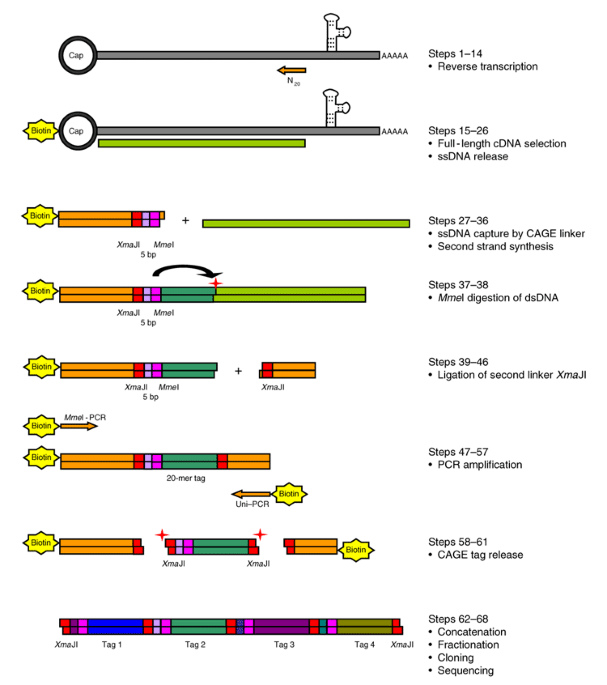
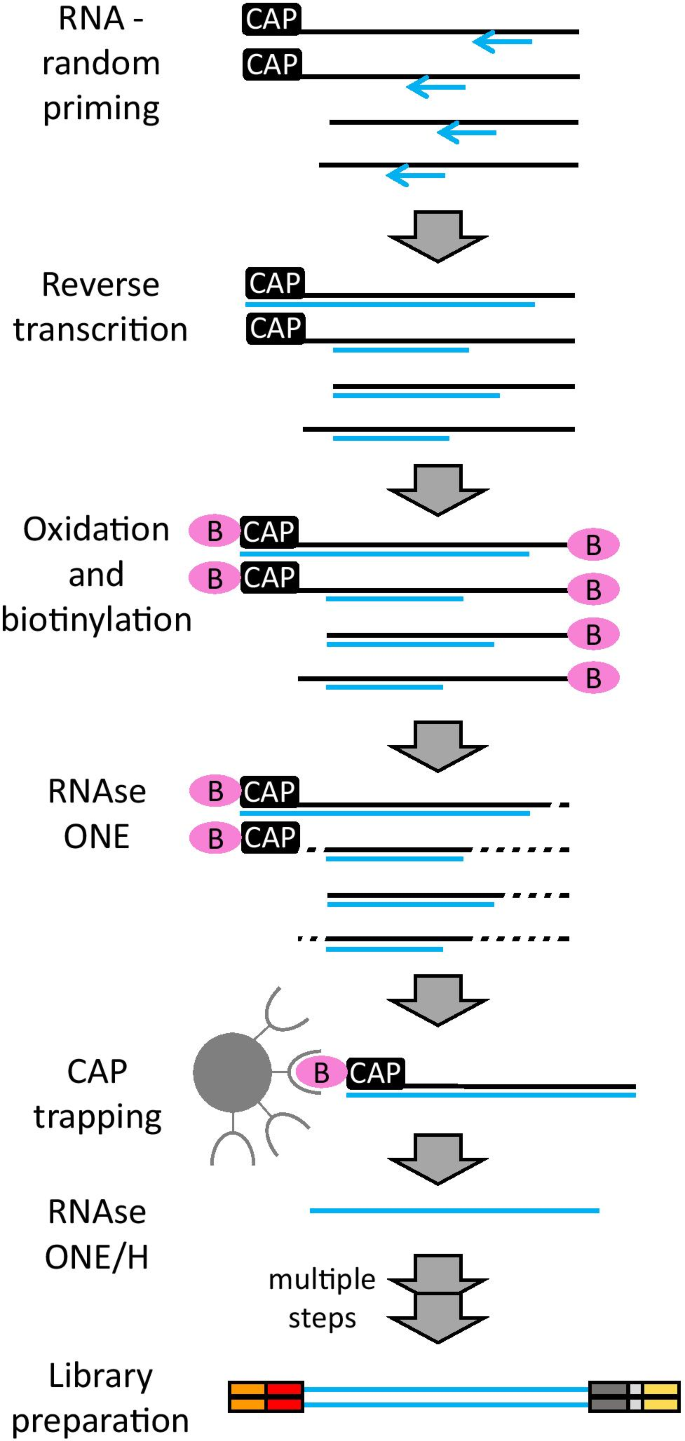
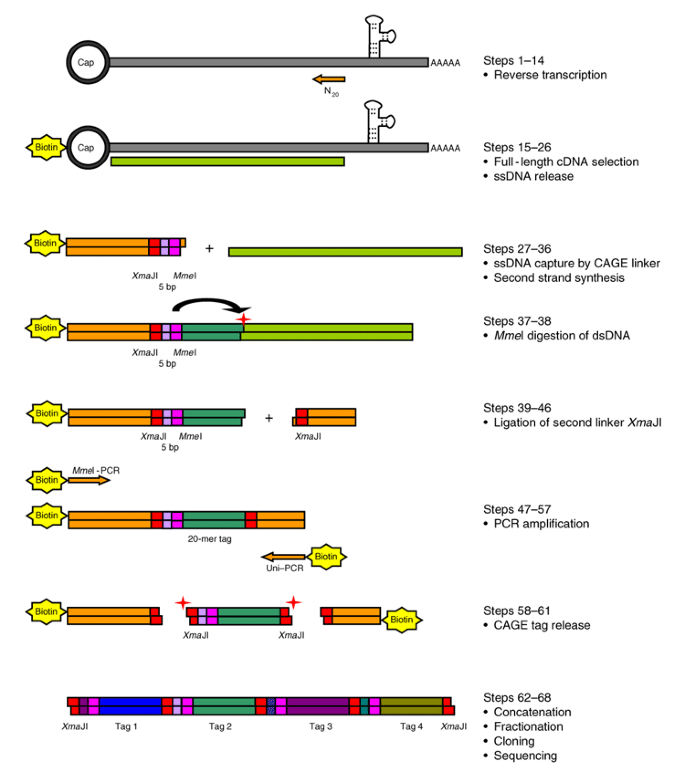
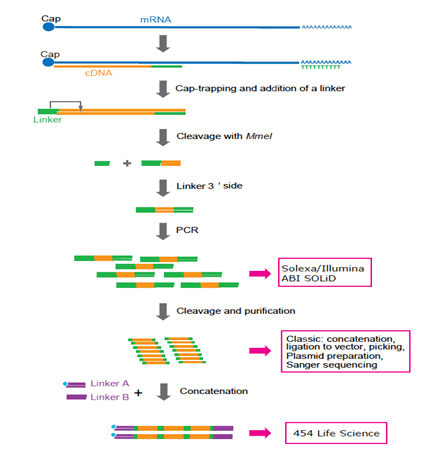
Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage | PNAS

CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology

CANONICAL ANALYSIS OF PRINCIPAL COORDINATES: A USEFUL METHOD OF CONSTRAINED ORDINATION FOR ECOLOGY - Anderson - 2003 - Ecology - Wiley Online Library

Canonical analysis of principal coordinates (CAP) ordination plot (Bray-Curtis) of fish assemblage data for each experimental treatment at each geographic location.